What is ClinVar?
For many people, getting their genome sequenced is the start of an exciting journey. Often, though, people are left unsure of where to begin their exploration of their DNA data. To learn how genetics affects health, a tool called ClinVar can be extremely useful. But, how exactly does this tool work? This tutorial will teach you!
The 99.9 percent
The human genome is made up of roughly 3 billion letters spread across 23 pairs of chromosomes. Together, these letters (also known as nucleotides) act as an instruction manual to determine our traits. From an individual’s hair color to their risk of heart disease, the genome contains the recipe that makes each person unique.
Randomly pick 2 people, and they likely have many more differences than similarities. One may be tall, need glasses, has a family history of breast cancer and hates the taste of cilantro. The other has red hair, is lactose intolerant, and has always been a straight-A student.
While these two people may seem nothing alike, they still share 99.9% of the same letters in their genomes. All of the variation between two random individuals, from their appearance to their disease risks, stems from differences in only 0.1% (or 3 million) of the letters in the genome!
The 0.1 percent
The letters of the genome that can be different between individuals are known as single nucleotide polymorphisms, or SNPs (pronounced “snips”). For example, a SNP is present if the majority of individuals have the letter “C” at a specific spot in the genome while a minority has a “T” instead. This SNP would be said to have 2 possible variants, or “alleles”: C and T.
Scientists estimate that there are potentially up to 100 million SNPs across the genome. Many of these SNPs occur within genes, the DNA sequences that act as instructions for making proteins, molecular machines that perform various tasks in and outside cells. Many more SNPs, though, occur outside of these regions, in the so-called “junk DNA”. Despite its name, this DNA comprises around 99% of our genome’s information, and scientists continue to discover more and more about its importance for our health.
Regardless, identifying SNPs is a key step in understanding personal genomic data. Some SNPs can have a powerful effect on an individual’s susceptibility to a certain disease. For example, consider Alzheimer’s disease. A particular SNP in the APOE gene increases a person’s risk of developing Alzheimer’s disease by more than 20-fold!
With advances in genetic testing technologies lowering the cost of sequencing a human genome, more and more people are taking the leap to learn about themselves through their DNA. Some genetic testing companies give their users access to their raw DNA files, to enable them to explore their data on their own. Though this empowers users, it can simultaneously be quite daunting. Most people are not very familiar with human genetics and various data analysis tools. In particular, given millions of SNPs, many users understandably don’t know how to learn what they mean.
Are you interested in discovering ALL genetic variants in your DNA to get most of tools like ClinVar? Nebula Genomics offers Whole Genome Sequencing for only $299! Click here to learn more!
How to use ClinVar
One of the most helpful tools to learn more about what your SNPs might mean for health is ClinVar. It is a freely accessible public archive that aims to catalog relationships between genetic variants and their impact on health status.
ClinVar is run by the National Institutes of Health in the United States. It works as a central, public database for researchers and medical professionals to deposit information about clinically relevant SNPs. For example, if a new scientific study is published linking a genetic variant to a particular disease, this information can be submitted to ClinVar.
Though ClinVar is powerful, it can oftentimes seem unwieldy and difficult to navigate. Learning how to effectively search through the site can unlock tons of additional information from your genome data.
At its core, you can search through ClinVar using any of 3 different pieces of information. These are:
- SNP ID
- Gene
- Disease/Condition
Searching ClinVar by SNP
Most SNPs are assigned a unique ID, generally starting with the letters “rs” and then a string of numbers (for example, rs7412). Entering this ID can directly connect you with information about the SNP’s location, any diseases it may be associated with, and much more.
Raw DNA files regardless of the format generally contain a list of genetic variants and their IDs. At Nebula Genomics, we provide our users with their genetic variants in the VCF format, which is the most commonly used format. To examine your genetic variants, you can VCF file with a text editor like Sublime or just take a look at the studies in the Nebula Library. To date, we have already analyzed over 8000 genetic variants, and new studies and variants are added every week!
To learn about the clinical relevance of a particular SNP let’s use ClinVar. Let’s start by navigating to the site, where the front page should look like this:
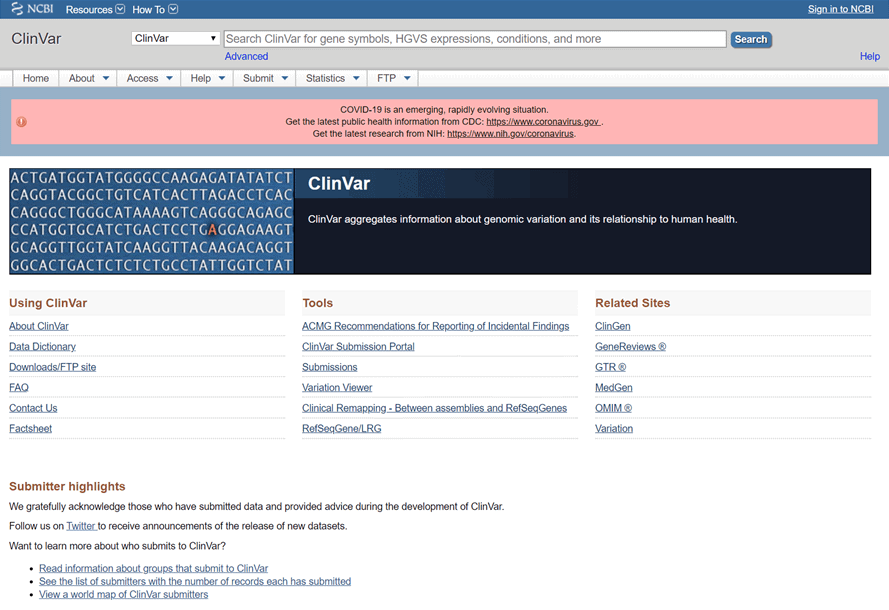
On this site, you can learn more about ClinVar and how to make new submissions. If you already have an SNP in mind, you can simply enter it into the search bar and press “Search”. Here, we are interested to learn more about SNP rs63750048.
If any information about a SNP has been uploaded to ClinVar, the site will return a search result for that SNP. We can see at a quick glance that it is located in a gene called PSEN2 and has been linked to Alzheimer’s disease type 4. Clicking on the link (highlighted in the screenshot), we can learn even more about the SNP.
On this page, more information about the SNP is provided, including all conditions it may be linked to, and any scientific publications that mention the SNP. For each condition in the “Submitted interpretations and evidence” section, you can click the “Evidence details” link to learn how the SNP is connected to the disease and how it was discovered.
Here, we can see the SNP was identified in an Italian family affected by Alzheimer’s disease. The normal letter at this location of the genome is C, but genetic tests performed on this family show many members have a T at the same location, which may affect how the PSEN2 gene functions. The report goes on to note specific symptoms that members of the family experienced.
Searching ClinVar by gene
Perhaps we do not have a particular SNP in mind, and would instead like to find all the SNPs present in a gene we’re interested in. Once we find an interesting SNP, we can return to our own genomic data and check our status for this SNP.
Start by navigating back to the ClinVar homepage, and search for the gene you are interested in learning more about. In this case, we search for “BRCA”, a gene where variations have been linked to an increased risk of developing breast cancer.
Searching this gene name returns hundreds of ClinVar variants. Some of them are in the BRCA genes and others are in genes closely related to BRCA. To avoid being overwhelmed by search results, we can filter them using options down the left side of the page. These include:
- Clinical significance: Does this SNP cause a particular disease (pathogenic and likely pathogenic) or not (benign and likely benign)? Or is it uncertain what its effects are? Or are there conflicting results from different studies?
- Molecular consequence: What effect does the SNP have on the gene’s sequence?
- Variation type: Is a letter added to the sequence or deleted from it? Or is a letter substituted to another (for example, C to T)?
- Review status: How much clinical evidence exists for this SNP?
After looking through the numerous SNPs at different filters, let’s learn more about the variant highlighted with the red box below.
By following the link, we can immediately learn more information about the SNP. We can see that the SNP is a change from the letter G to the letter C located on chromosome 13, and it is potentially linked to an increased risk of breast cancer. Additionally, we can learn the ID of the SNP (rs81002796).
Searching ClinVar by disease
Finally, if we don’t have a particular SNP or a particular gene in mind, we can search through all the SNPs linked to a particular disease. Though not required, adding the [disease/phenotype] tag after a search term can help limit results to only the most pertinent for the specific condition. For example, if we are interested in SNPs connected to diabetes, we can search ClinVar with the term “diabetes[disease/phenotype]”.
Since thousands of results are returned, let’s filter out the results to only display those that are established “risk factors” for diabetes. Then, let’s learn more about the first result.
By following the link, we can learn that this SNP is the change of a letter C to G located on chromosome 1 in a gene known as PTPN22. The SNP was identified in a publication in 2006.
If we are interested in learning more about how the SNP was found, we can follow the publication’s link to learn more about the study.
ClinVar is powerful. Use it wisely.
Though ClinVar is an expansive database compiling results from thousands of laboratory and medical studies, it is important to note that the tool is not a genetic counselor. It is not designed to provide medical advice and diagnoses. If you suspect that the information you have found throughout your searches on the site might be relevant to your health, it is important to see a medical doctor or another licensed health care professional for genetic counseling.
Did you like our ClinVar tutorial? You might find these tutorial useful as well:
- How to download your 23andMe and AncestryDNA data?
- How to use GEDmatch?
- How to use the Nebula Library?
- How to explore your genomic data?
- How does paternity testing work?
- DNA testing during pregnancy?
