Edited by Christina Swords, Ph.D.
Genetic testing has never been easier. At-home DNA testing services continue to grow in popularity as ways to learn more about your ancestry and inherited traits. However, sifting through all of the companies and DNA testing types to find the best DNA test can be overwhelming. Here, we give you a brief overview of the different types of DNA tests.
DNA Testing Technologies
DNA microarrays and Next Generation Sequencing (NGS) are two different technologies commonly used for genetic testing. Both can be used to read out genetic information from chromosomal and mitochondrial DNA. However, DNA microarrays and NGS technologies work very differently.
DNA Microarray-based Genotyping
DNA microarrays read out DNA at a defined set of positions that are known to vary between people. These positions are called Single Nucleotide Polymorphisms (SNPs). Most genetic testing services, such as 23andMe and AncestryDNA, use DNA microarray technology to profile (genotype) your genome at ~ 500,000 positions. This is less than 0.1% of the whole human genome. While microarray-based genotyping is very affordable, it misses a lot of important information.
Next Generation Sequencing
Next Generation Sequencing (NGS) is a technology that enables reading large amounts of genetic information very efficiently. NGS works by reading many small stretches of DNA and then piecing them together to determine a continuous sequence (Figure 1). In contrast to DNA microarrays, NGS can be used to read out entire genomes instead of just a small number of positions. However, while NGS produces much more data than microarray-based genotyping it tends to be more expensive. There exist several NGS-based DNA tests that have different price points.
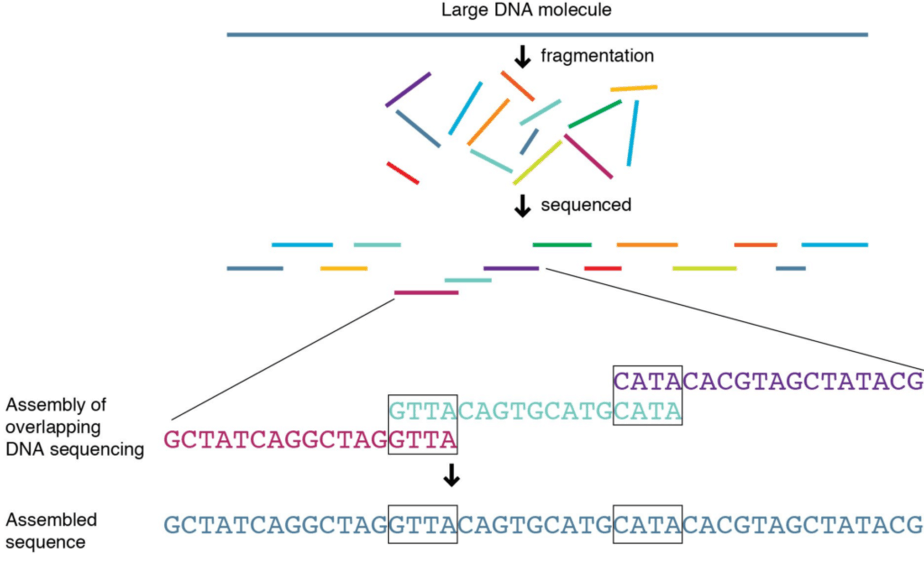
Figure 1. Next Generation Sequencing (NGS). Image Courtesy of National Human Genome Research Institute.
NGS-based DNA Tests
Whole Exome Sequencing (WES)
Whole Exome Sequencing (WES) reads out the ~ 1.5% of the human genome that encode proteins – molecular machines that perform most of the cellular functions. Because WES covers only a small portion of a human genome it is cheaper than WGS. However, WES misses genetic variants that are outside protein-coding regions and often have important regulatory functions. Furthermore, Whole Exome Sequencing is still quite expensive when compared to microarray-based genotyping. Thus, our answer to the question of whether to choose Whole Genome Sequencing vs Whole Exome Sequencing is to go with Whole Genome Sequencing.
Whole Genome Sequencing (WGS)
NGS technology can be used to read out an entire human genome. This is referred to as Whole Genome Sequencing (WGS). It enables unbiased, comprehensive discovery of genetic variants and yields deeper insight into personal genetic makeup. In addition to reading the entire genome, each position is often read many times in order to increase result accuracy. For example, WGS is typically done at 30x coverage (depth) which means that on average each position in the genome is read 30 times. This makes WGS much more expensive than microarray-based genotyping.
Nebula Genomics
Our mission at Nebula Genomics is to make personal genome sequencing affordable to everyone. In February 2020, we achieved an important milestone by bringing the cost of 30x Whole-Genome Sequencing below $300. This brings the cost of personal genome sequencing close to the cost of much less comprehensive genetic tests.
Do you want to learn more about how DNA testing technologies are applied? Take a look at our introduction to paternity testing!