Imagine your genome like an orchestra. Sometimes, sound flows from just one instrument, a gifted soloist that single-handedly fills the concert hall. But more often, the music is an amalgamation of many different instruments, each one making a small contribution to the overall tune. Today, genetic research seeks to uncover how an orchestra of genes determines various traits and how diseases occur if that orchestra is out of tune.
The genome’s roughly 20,000 genes work in a similar way. A particular trait or disease can be the work of a single gene — for example, blood type and cystic fibrosis. But the most common diseases and traits, such as heart disease, diabetes, height, and intelligence, are the result of multiple genes functioning in combination. In fact, hundreds, even thousands of genes can work together, each one exerting a slight influence on human biology. Today, scientists are conducting genetic research to better dissect these polygenic traits — that is, which genes and genetic variations (mostly single nucleotide polymorphisms) contribute and how much?
Nebula Library – Genetic Research Unlocked
At Nebula Genomics, we are committed to bringing cutting-edge scientific discoveries to our users, as well as making their meaning more intuitive to grasp. We are excited to announce the launch of the Nebula Research Library – a weekly updated collection of personalized reports based on the latest genetic discoveries. The Nebula Research Library is the core part of our reporting. With it, you will be able to stay up to date with the latest discoveries in human genomics and how they may relate to you. Today, the Nebula Library includes over 150 genome-wide association studies (GWAS). We made the content of the Nebula Library publicly available on our blog to facilitate exploration and discovery of content that interests you.
In this blog post, we’ll explain how to use the Nebula Research Library and understand the results of the genome-wide association studies that it contains.
Genetic Research Explained
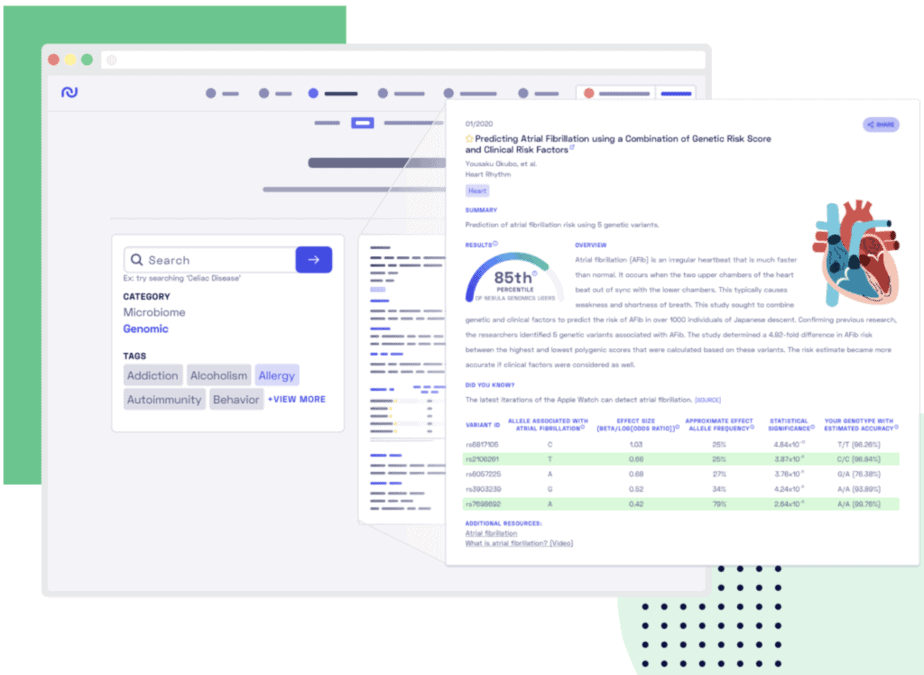
Figure 1 shows an exemplary library entry – a genome-wide association study that examined the genetics of attention deficit hyperactivity disorder (ADHD). First, you are presented with the title of a research article, along with publication date, lead author and name and publication journal. To the right of the article title, you can find a share button that you can use to share your results over email or on social media. Below the title, there are topic tags that will help you explore the Nebula Research Library for related content.
Next, we provide a summary of the study along with some additional information that is relevant to the condition studied. In the study example below we explain what ADHD is, provide some information about how the study was conducted. For example, the number of study participants and their ethnicity.
Polygenic Scores
We calculate two numbers for most of the studies. The first number is a polygenic score. It summarizes the effect of all genetic variants discovered in the study and present in your genome. The second number is a percentile that is calculated for the polygenic score. It tells you how your polygenic score compares to the scores of other Nebula Genomics users.
Figure 1 shows a polygenic score for attention deficit hyperactivity disorder (ADHD) that was calculated from ADHD-associated variants identified in this study (A). This score places the user into the 97th percentile (B), which means that he has a higher polygenic score for ADHD than 97% of our users.
Figure 1. An entry in the new Nebula Research Library.
Detailed Genetic Information
Below the polygenic score, you can find a table with genetic variants. The first two columns of the table contain the variant IDs and the alleles – versions of a variant – that have been found to be associated with a trait or disease, which is ADHD in the example in Figure 1 (C).
Next, come three columns that list allele effect sizes, frequencies and statistical significance of the associations (D). The effect sizes are the contributions of different alleles to the polygenic score. Some alleles have positive effect sizes and increase the score (highlighted in green) while other alleles have negative effect sizes and decrease the score (highlighted in blue). Alleles not present in your genome do not affect your polygenic score (not highlighted). The next column contains allele frequencies which tell you the percentage of people in the population that carry each of the listed alleles. We also show statistical significance for each of the associations discovered in a study. These numbers are called p-values. The smaller the p-value, the more certain it is that the discovered association between a trait and a genetic variant is real.
The last column shows your genotype – the two alleles you have inherited from your parents – and an estimated accuracy of our prediction (E). Note that we account for this uncertainty when calculating your polygenic scores. If the accuracy is low, then the contribution to the polygenic score is reduced.
Proceed with Caution
We would like to emphasize that the scores that we calculate should not be considered genetic risk scores and not used for predicting the risk of developing diseases. There are several limitations.
First, polygenic scores describe only genetic predispositions. The influences of environmental factors such as lifestyle are not considered. Second, the genetic heritability of most traits and diseases remains poorly understood. The studies in the Nebula Library list only the most significant associations, which are only a small fraction of all existing links between genetics and traits. The identified variants are also not necessarily the causal variants. Third, most studies examined genetic data from individuals of European ancestry. Whether the same associations exist in other populations and can be used for risk prediction often remains an open question. Large population genomics projects like the UK Biobank are underway to answer this question.
For these reasons, the Nebula Research Library is not intended for medical or diagnostic use. Genes are not necessarily destiny. If you have a particular variant associated with a disease in your genome, it does not mean you will develop that disease. The same goes for polygenic scores. A high polygenic score/percentile for a particular disease does not necessarily mean a significantly increased disease risk. As always, if you have any questions about your health, please seek the advice and input of a healthcare provider.
Join Us Today
Personal genome sequencing has become affordable to almost everyone. You can now enter the age of personal genomics and learn about how the latest discoveries relate to your DNA. Order our whole-genome sequencing or get started for free by uploading your 23andMe/AncestryDNA data!
Reports in the Nebula Library (August 2020)
- Multiple Sclerosis (IMSGC, 2019)
- Parental Lifespan (Timmers, 2020)
- Longevity (Timmers, 2020)
- Healthspan (Timmers, 2020)
- Gambling disorders (Lind, 2012)
- Inflammatory protein level (Hillary, 2020)
- Tourette’s syndrome (Yu, 2019)
- Anxiety (Meier, 2019)
- Autoimmune thyroid disease (Saevarsdottir, 2020)
- Chronic wound microbiome diversity (Tipton, 2020)
- Brain lesions (Armstrong, 2020)
- Corneal thickness (Choquet, 2020)
- Type 2 diabetes (Vujikovic, 2020)
- Thinness (Orthofer, 2020)
- Long QT Syndrome (Lahrouchi, 2020)
- Triple-negative breast cancer (Zhang, 2020)
- HER2-enriched-like breast cancer (Zhang, 2020)
- Luminal B-like breast cancer (Zhang, 2020)
- Luminal B/HER2-negative-like breast cancer (Zhang, 2020)
- Luminal A-like breast cancer (Zhang, 2020)
- PR interval (Ntalla, 2020)
- Carbohydrate consumption (Meddens, 2020)
- Fat consumption (Meddens, 2020)
- Protein consumption (Meddens, 2020)
- Sugar consumption (Meddens, 2020)
- Left ventricular ejection fraction (Pirruccello, 2020)
- Stroke volume (Pirruccello, 2020)
- Left ventricular end-systolic volume (Pirruccello, 2020)
- Left ventricular end-diastolic volume (Pirruccello, 2020)
- Type 2 diabetes (Spracklen, 2020)
- Melanoma (Landi, 2020)
- Lung cancer (McKay, 2017)
- Squamous cell lung carcinoma (McKay, 2017)
- Lung adenocarcinoma (McKay, 2017)
- Sjögren’s syndrome (Lessard, 2013)
- Computer use for leisure (van de Vegte, 2020)
- Driving for leisure (van de Vegte, 2020)
- Television watching for leisure (van de Vegte, 2020)
- Gallstones (Ferkingstad, 2018)
- Atopic dermatitis (Paternoster, 2015)
- QT interval (Arking, 2014)
- Moles (Falchi, 2009)
- Age-related macular degeneration (Han, 2020)
- Hydroxyvitamin D level (Revez, 2020)
- Membranous nephropathy (Xie, 2020)
- Febrile seizures (Feenstra, 2014)
- Total Brain Volume (Zhao, 2019)
- Parkinson’s Disease (Nalls, 2019)
- Subcortical Brain Volume (Satizabal, 2019)
- Well-Being (Baselmans, 2019)
- Allergic Rhinitis (Waage, 2018)
- Birth Weight and Disease Susceptibility (Horikoshi, 2017)
- Loneliness (Abdellaoui, 2019)
- Albuminuria (Teumer, 2019)
- Aortic Calcification (Malhotra, 2019)
- Venous Thromboembolism (Klarin, 2019)
- Systemic Sclerosis (López-Isac, 2019)
- Tremors (Müller, 2016)
- Body Mass Index (Couto Alves, 2019)
- Reproductive Behavior (Barban, 2016)
- Diffuse Lung B-Cell Lymphoma (Kleinstern, 2019)
- Addiction (Food) (Cornelis, 2016)
- Eosinophilic granulomatosis with polyangiitis (Lyons, 2019)
- Bone Density (Kemp, 2017)
- Gastroesophageal Reflux (An, 2019)
- Alzheimer’s Disease (Kunkle, 2019)
- Hearing Loss (Wells, 2019)
- Healthy Lung Function (Shrine, 2019)
- Accelerated Aging (Gibson, 2019)
- Neuroticism (Luciano, 2017)
- Loss of Chromosome Y (Thompson, 2019)
- Fainting (Hadji-Turdeghal, 2019)
- Brain age (Jonsson, 2019)
- Acute lymphoblastic leukemia (Vijayakrishnan, 2019)
- Varicose Veins (Fukaya, 2018)
- Schizophrenia (Lam, 2019)
- Age-Related Macular Degeneration (Fritsche, 2015)
- Brugada Syndrome (Bezzina, 2013)
- Gout (Tin, 2019)
- Inflammation Markers in the Blood (Ligthart, 2018)
- Chronic Lymphocytic Leukemia (Berndt, 2016)
- Psychosis (Legge, 2019)
- Glaucoma (MacGregor, 2018)
- Mosaic Loss of Chromosome Y (Terao, 2019)
- Refractive errors (Hysi, 2020)
- Neck or shoulder pain (Meng, 2020)
- Male puberty timing (Hollis, 2020)
- Resting Heart Rate (Eppinga, 2016)
- Optic Disc Size (Hun, 2019)
- Type 2 diabetes (Cook, 2016)
- Attractiveness to Mosquitoes (Jones, 2017)
- Retinal Detachment (Boutin, 2019)
- Thinning Cornea: Keratoconus (McComish, 2019)
- Anhedonia, an inability to feel pleasure (Ward, 2019)
- Malaria Resistance (Malaria Genomic Epidemiology Network, 2019)
- Nasal Polyps (Kristjansson, 2019)
- Amyotrophic Lateral Sclerosis (Nicolas, 2019)
- Helicobacter pylori Infection (Mayerle, 2013)
- Kawasaki Disease (Onouchi, 2012)
- Atrial Fibrillation (Okubo, 2020)
- Heavy Alcohol Consumption (Thompson, 2020)
- Height (Wood, 2014)
- Facial Attractiveness (Hu, 2019)
- Breast Cancer (Fachal, 2020)
- Heart Failure (Shah, 2020)
- Diet and Schizophrenia (Niarchou, 2020)
- Diverticular Disease (Maguire, 2018)
- Lupus (Chen, 2020)
- Fraternal (dizygotic) Twins (Mbarek, 2016)
- Dietary Habits (Matoba, 2020)
- Respiratory distress syndrome (Guillen-Guio, 2020)
- Glaucoma (Craig, 2020)
- Hemochromatosis (Benyamin, 2014)
- Cutaneous Squamous Cell Carcinoma (Sarin, 2020)
- Snoring (Campos, 2020)
- Prion Disease (Mead, 2009)
- Coffee Consumption (Zhong, 2019)
- Telomere Length (Li, 2020)
- Opioid use (Polimanti, 2020)
- Crohn’s disease (Franke, 2010)
- Idiopathic pulmonary fibrosis (Allen, 2019)
- Alopecia areata (Petukhova, 2010)
- Podoconiosis (Ayele, 2012)
- Body Mass Index (BMI) (Anderson, 2020)
- Celiac Disease (Dubois, 2010)
- Testosterone (Ruth, 2020)
- HIV Resistance (Samson, 1996)
- Vitamin D (Manousaki, 2020)
- Aplastic Anemia (Savage, 2020)
- Telomere Length (Codd, 2013)
- Male-Pattern Baldness (Pirastu, 2017)
- Knee Pain (Meng, 2019)
- Blood Cancer (Lin, 2019)
- Genetics and Income (Hill, 2019)
- Vitiligo (Jin, 2017)
- Triglyceride level (Richardson, 2020)
- LDL cholesterol level (Richardson, 2020)
- HDL cholesterol level (Richardson, 2020)
- Apolipoprotein B level (Richardson, 2020)
- Apolipoprotein A-1 level (Richardson, 2020)
- Migraine (Anttila, 2013)
- Handedness (Wiberg, 2009)
- Visceral Adiposity (Karlsson, 2019)
- Gestational Duration (Liu, 2019)
- Chronic Kidney Disease (Hellwege, 2019)
- Cerebral Small Vessel Disease (Chung, 2019)
- Autism Spectrum Disorder (Grove, 2016)
- Blood Pressure (Warren, 2019)
- Rheumatoid Arthritis (Stahl, 2011)
- Cerebral cortex thickness (Grasby, 2020)
- Familial short stature (Lin, 2020)
- Pelvic organ prolapse (Olafsdottir, 2020)
- Cerebral cortex surface area (Grasby, 2020)
- Breast cancer (Shu, 2020)
- Longevity (Deelen, 2019)
- Thyroid cancer (Liyanarachchi, 2020)
- SARS coronavirus infection (Ching, 2010)
- SARS coronavirus infection (Hamano, 2005)
- SARS coronavirus infection (Tu, 2015)
- Subcortical Brain Structures (Hilbar, 2015)
- Anorexia Nervosa (Watson, 2019)
- Asthma (Johansson, 2019)
- Colorectal Cancer (Huyghe, 2018)
- Depression (Wray, 2018)
- Alzheimer’s Disease (Wang, 2019)
- Intelligence (Sniekers, 2017)
- Alzheimer’s Disease (Deming, 2019)
- Cholesterol Levels (Kathiresan, 2008)
- Ulcerative Colitis (Anderson, 2011)
- Venous Thromboembolism (Lindstrom, 2019)
- Peripheral Artery Disease (Klarin, 2019)
- Personality (Lo, 2017)
- Alcohol Consumption (Evangelou, 2019)
- Risky Behavior (Linnér, 2019)
- PTSD (Gelernter, 2019)
- Acne (Navarini, 2019)
- Kidney Stones (Thorleifsson, 2009)
- Menopause (Stolk, 2012)
- Migraine (Freilinger, 2012)
- Melanoma (Barrett, 2011)
- Depression (Power, 2017)
- Blood Pressure (Surendran, 2016)
- Ulcerative Colitis (UK IBD Genetics Consortium, 2009)
- Alzheimer’s Disease (Seshadri, 2010)
- Restless Leg Syndrome (Winkelmann, 2011)
- Chronic Kidney Disease (Chambers, 2010)
- PTSD (Logue, 2012)
- Coronary Artery Disease (Lee, 2013)
- Stroke (Malik, 2018)
- Obesity (Jiao, 2011)
- Cannabis Use Disorder (Demontis, 2019)
- Cleft Lip (Yun, 2019)
- Psoriasis (Tsoi, 2015)
- Asthma (Ferreira, 2019)
- Daytime Sleepiness (Wang, 2019)
- Primary Biliary Cirrhosis (Mells, 2011)
- Alcohol Consumption (Jorgenson, 2017)
- Bladder Cancer (Figueroa, 2013)
- Breast Cancer (Michailidou, 2013)
- Schizophrenia (Psychiatric GWAS Consortium, 2011)
- Bipolar Disorder (Mühleisen, 2014)
- Insomnia (Hammerschlag, 2017)
- Thyroid Cancer (Gudmundsson, 2017)
- Tuberculosis (Thye, 2012)
- Pancreatic Cancer (Childs, 2015)
- Prostate Cancer (Schumacher, 2011)
- Epilepsy (ILAEC, 2014)
- Insomnia (Lane, 2016)
- Osteoarthritis (arcOGEN Consortium, 2012)
- Adolescent idiopathic scoliosis (Kou, 2019)
- Attention-deficit/Hyperactivity Disorder (Demontis, 2018)
- Glioma (Shete, 2009)
- Emphysema (Parker, 2019)